Old ARC Cluster
Looking for the documentation of the old ARC
Cluster, or the even older ARC Cluster?
Hardware Status
- c[63-66] openmpi3 is not working.
- c[30-57] have been retired.

READ THIS BEFORE YOU RUN ANYTHING: Running Jobs (via Slurm)
(1) Notice: Store large data files in the Beegfs file
system and not your home directory. The home directory is for
programs and small data files, which should not exceed 10GB altogether.
(2) Once logged into ARC, immediately obtain access to a compute node
(interactively) or schedule batch jobs as shown below. Do not
execute any other commands on the login node!
- interactively:
- srun -n 16 --pty /bin/bash # get 16 cores (1 node) in interactive mode
- mpicc -O3 /opt/ohpc/pub/examples/mpi/hello.c # compile MPI program
- prun ./a.out # execute an MPI program over all allocated nodes/cores
- in batch mode:
- compile programs interactively beforehand (see above)
- cp /opt/ohpc/pub/examples/slurm/job.mpi . # script for job
- cat /opt/ohpc/pub/examples/slurm/job.mpi # have a look at it, executes a.out
- sbatch job.mpi # submit the job, wait for it to get done:
creates job.%j.out file, where %j is the job number
- more slurm options:
- srun -n 32 -X --pty /bin/bash # get 32 cores (2 nodes) in interactive mode with X11 graphical output
- srun -n 16 -N 1 --pty /bin/bash # get 1 interactive node with 16 cores
- srun -n 32 -N 2 -w c[90,91] --pty /bin/bash #run on nodes 90+91
- srun -n 64 -N 4 -w c[90-93] --pty /bin/bash #run on nodes 90-93
- srun -n 64 -N 4 -p broadwell --pty /bin/bash #run on any 4 broadwell nodes
- srun -n 64 -N 4 -p rtx2070 --pty /bin/bash #run on any 4 nodes with RTX 2070 GPUs
- sinfo #available nodes in various queues, queues are listed in "Hardware" section
- squeue # queued jobs
- scontrol show job=16 # show details for job 16
- scancel 16 # cancel job 16
- Slurm documentation
- Slurm
command summary
- Editors are available once on a compute node:
- inside a terminal: vi, vim, emacs -nw
- using a separate window: evim, emacs

Hardware
1280 cores on 80 compute nodes integrated by
Advanced HPC. All machines are
2-way SMPs (except for single socket AMD Rome/Milan machines) with either
AMD or Intel processors (see below) and a total 16 physical cores per node (32 for c[58-59]).
Nodes:
- AMD Opteron: nodes c[60], queue: -p opteron
- Intel Sandy Bridge: nodes c[95-98], queue: -p sandy
- Intel Ivy Bridge: nodes c[99-100], queue: -p ivy
- Intel Broadwell: nodes c[78-94], queue: -p broadwell
- Intel Skylake Silver: nodes c[0-19,26-29], queue: -p skylake
- AMD Epyc Rome: nodes c[20-25,61,65-77,101-107], queue: -p rome
- Intel Cascade Lake: nodes c[63-64], queue: -p cascade
- AMD Epyc Milan: nodes c[58-59,62], queue: -p milan
- login node: arcl (1TB
HDD+Intel
X520-DA2 PCI Express 2.0 Network Adapter E10G42BTDABLK)
- nodes: cXXX, XXX=0..107 (1TB HDD)
- 16 nodes with NVIDIA Quadro P4000 (8 GB, sm 6.1): nodes c[0-3,8-19], queue: -p p4000
- 12 nodes with NVIDIA RTX 2060 (6 GB, sm 7.5): nodes c[26,29,79-85,88-90], queue: -p rtx2060
- 15 nodes with NVIDIA RTX 2070 (8 GB, sm 7.5): nodes c[78,91-95,97-104,107], queue: -p rtx2070
- 2 nodes with NVIDIA RTX 2080 (8 GB, sm 7.5): nodes c[24,28], queue: -p rtx2080
- 22 nodes with NVIDIA RTX 2060 Super (8 GB, sm 7.5): nodes c[20-21,25,60,62,65-77,86-87,96,106], queue: -p rtx2060super
- 2 nodes with NVIDIA RTX 2080 Super (8 GB, sm 7.5): nodes c[22-23], queue: -p rtx2080super
- 2 nodes with NVIDIA RTX 3060 Ti (8 GB, sm 8.6): node c[27,105], queue: -p rtx3060ti
- 6 nodes with NVIDIA RTX A4000 (16 GB, sm 8.6): node c[4-7,63-64], queue: -p a4000
- 2 nodes with 4 GPUs each NVIDIA RTX A6000 (48 GB, sm 8.6): node c[58-59], queue: -p a6000
- 1 node with NVIDIA A100 (80 GB, sm 8.6): node c[61], queue: -p a100
- 2 nodes with NVIDIA RTX 4060 Ti (8 GB, sm 8.7): node c[26,29], queue: -p rtx4060ti
- Altera Arria 10 FPGA on c82
- Altera Stratix 10 FPGA on c27,c28
- head node: arch (has 8TB SSD RAID5 using 10xSamsung
960 PM863a) with
Supermicro X10DRU-i Motherboard and
Transport
SYS-1028U-TRT plus a
10GEther
4xSPF+ Broadcom BCM57840S card 20Gbps bonded (dynamic link aggregation) to internal GEther switches
- backup node: arcb (same configuration as arch, except no 10GEther card)





















|
Networking, Power and Cooling:
Pictures

System Status



Software
All software is 64 bit unless marked otherwise.

Obtaining an Account
- for NCSU students/faculty/staff in Computer Science:
- Send an email to your advisor asking for ARC access and indicate your unity ID.
- Have your advisor endorse and forward the email
to Subhendu Behera.
- If approved, you will be sent a secure link to upload your
public RSA key (with a 4096 key length) for SSH access.
- for NCSU students/faculty/staff outside of Computer Science:
- Send a 1-paragraph project description with estimated compute
requirements (number of processors and compute hours per job per
week) in an email to your advisor asking for ARC access and indicate your unity ID.
- Have your advisor endorse and forward the email
to Subhendu Behera.
- If approved, you will be sent a secure link to upload your
public RSA key (with a 4096 key length) for SSH access.
- for non-NCSU users:
- Send a 1-paragraph project description with estimated compute
requirements (number of processors and compute hours per job per
week) in an email to your advisor asking for ARC access. Indicate
the hostname and domain name that you will login from (e.g., sys99.csc.ncsu.edu).
- Have your advisor endorse and forward the email
to Utsab Ray.
- If approved, you will be sent a secure link to upload your
public RSA key (with a 4096 key length) for SSH access.

Accessing the Cluster
- Login for NCSU users:
- Login to a machine in the .ncsu.edu domain (or use NCSU's VPN).
- Then issue:
- Or use your favorite ssh client under Windows from an .ncsu.edu
machine.
- Login for users outside of NCSU:
- Login to the machine that your public key was generated on.
Non-NCSU access will only work for IP numbers that have been
added as firewall exceptions, so please use only the computer
(IP) you indicated to us any other computer will not work.
- Then issue:
- Or use your favorite ssh client under Windows.

Using OpenMP (via gcc/g++/gfortran)
-
The "#pragma omp" directive in C/C++ programs works.
gcc -fopenmp -o fn fn.c
g++ -fopenmp -o fn fn.cpp
gfortran -fopenmp -o fn fn.f
-
To run under MVAPICH2 on Opteron nodes (4 NUMA domains over 16 cores),
it's best to use 4 MPI tasks per node, each with 4 OpenMP threads:
export OMP_PROC_BIND="true"
export OMP_NUM_THREADS=4
export MV2_ENABLE_AFFINITY=0
unset GOMP_CPU_AFFINITY
mpirun -bind-to numa ...
-
To run under MVAPICH2 on Sandy/Ivy/Broadwell nodes (2 NUMA domains over 16 cores),
it's best to use 2 MPI tasks per node, each with 8 OpenMP threads:
export OMP_PROC_BIND="true"
export OMP_NUM_THREADS=8
export MV2_ENABLE_AFFINITY=0
unset GOMP_CPU_AFFINITY
mpirun -bind-to numa ...

Running CUDA Programs (Versions 8.0, 10.0, 11.X)

Running MPI Programs with MVAPICH2 and gcc/g++/gfortran (Default)

Running MPI Programs with Open MPI and gcc/g++/gfortran (Alternative)
- Issue
module switch mvapich2 openmpi
or, for new version,
module switch gnu gnu8
module switch mvapich2 openmpi3
-
Compile MPI programs:
mpicc -O3 -o pi pi.c
mpic++ -O3 -o pi pi.cpp
mpifort -O3 -o pi pi.f
-
Execute the program on 2 processors (using Open MPI):
prun ./pi
-
Execute the program on 32 (virtual) processors using 16 (physical)
cores (using Open MPI):
mpirun --oversubscribe -np 32 ./pi
- switch back to MVAPICH2:
module switch openmpi mvapich2
or, for new version,
module switch gnu8 gnu
module switch openmpi3 mvapich2
Using the PGI compilers (V16.7 for CUDA 8.0 capable GPU nodes, V19.10 for all others)
(includes OpenMP and CUDA support via pragmas, even for Fortran)
- Issue
module unload cuda
module load pgi
- For Fortran 77, use: pgf77 -V x.f
- For Fortran 95, use: pgf95 -V x.f
- For HPF, use: pghpf -V x.f
- For C++, use: pgCC -V x.c
- For ANSI C, use: pgcc -V x.c
- For debugging, use: pgdbg
- For more compile output, add option: -Minfo=all
- For AMD 64-bit, add option: -tp=barcelona-64
- For OpenMP, add option: -mp
- For OpenACC/CUDA (default: 7.0), add options: -acc
- For OpenACC/CUDA 7.5: -acc -Mcuda=cuda7.5,rdc
- For OpenACC/CUDA 8.0: -acc -Mcuda=cuda8.0,rdc
- For OpenACC/CUDA on specific GPUs, run: pgaccelinfo, then use the respective -ta option in the output for compilation, e.g., -acc -ta=tesla,cc20
- with filename.f: supports Fortran ACC pragmas (for CUDA), e.g.,
!$acc parallel
- with filename.c: supports C ACC pragmas (for CUDA), e.g.,
#pragma acc parallel
- Slides and
excercises on MPI+GPU programming with CUDA and OpenACC
- PGI Documentation
- OpenAcc Documentation
- PGI+Open MPI issue:
module switch mvapich2 openmpi/1.10.4
export LMOD_FAMILY_MPI=openmpi
#compile for C (similar for C++/Fortran)
mpicc ...
#use mpirun or prun to execute
prun ./a.out
mpirun ./a.out
Notice that this OpenMPI version has support for
CUDA pointers,
RDMA, and GPU Direct
- PGI+MVAPICH2: not supported

Dynamic Voltage and Frequency Scaling (DVFS)
- Change the frequency/voltage of a core to save energy (without
any of with minor loss of performance, depending on how memory-bound
an application is)
- Use cpupower
and its
utilities
to change processor frequencies
- Example for core 0 (requires sudo rights):
- cpupower frequency-info
- sudo cpupower -c 0 frequency-set -f 1200Mhz #set to userspace 1.2GHz
- cpupower frequency-info
- sudo cpupower frequency-set -g ondemand #revert to original settings
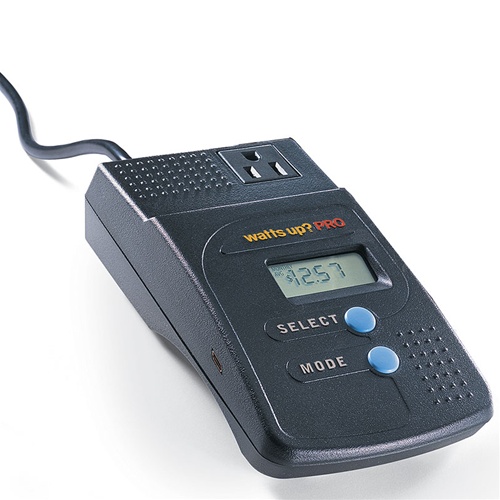

Power monitoring
Sets of three compute nodes share a power meter; in such a set,
the lowest numbered node has the meter attached (either on the serial
port or via USB). In addition, two individual compute nodes have power
meters (with different GPUs). See
this power wiring diagram to identify
which nodes belong to a set. The diagram also indicates if a meter
uses serial or USB for a given node. We recommend to explicitly
request a reservation for all nodes in a monitored set (see srun
commands with host name option). Monitoring at 1Hz is accomplished
with the following software tools (on the respective nodes where
meters are attached):
Virtualization with LXD (optionally with X11, VirtualBox, Docker inside)
Container virtualization support is realized
via LXD. Please try to
use CentOS images as they will take much less space than any other
ones since only the differences to the host image need to be stored
in the container. Also, do NOT deploy LXD on nodes c[0-19] as
they host BeeGFS. LXD/docker has been known to lock up nodes, and if
this happens on nodes c[0-19], it would affect other users on other
nodes as the BeeGFS file system would not longer be
operational. Finally, stop and delete images before you release
a node reserved by srun!
- lxd init #press enter to select defaults/empty password
(encouraged), or choose specific settings (discouraged)
If this does not work, send us email (see above), lxd is sometimes
problematic in its setup.
- lxc image list images:|grep -i centos #list of centos images
- lxc launch images:centos/7/amd64 my-centos #create and start new image
- lxc list #see installed/running images
- lxc exec my-centos -- /bin/bash #get a shell for running image
- yum install openssh-server
- systemctl start sshd
- passwd #enter root passwords
- #install other useful packges (see CentOS 7 docs), e.g., gcc compiler:
- yum group install "Development Tools"
- #from login node, create another session to your compute node, say cXX:
- ssh cXX
- #using the IP from "lcx list", transfer files over the virtual bridge to lxc image:
- scp some-file root@10.196.17.XXX: #or use sftp
- lxc stop my-centos #stop the image
- lxc config device add my-centos gpu gpu #optionally add GPU
support, then you need to install CUDA
- lxc start my-centos #start the image
- lxc delete my-centos #delete all files of the image
- Further
instructions
for Ubuntu, skip install steps and just look at user commands (lxc)
Notice: Images are installed locally on the node you are
running on. If you need identical images on multiple nodes, then write
a script to create an image from scratch. You cannot simply copy
images as they are in a protected directory.
X11 inside LXD:
- lxc exec my-centos -- /bin/bash
- yum -y install openssh-server xauth xeyes
- systemctl start sshd
- useradd myuser
- passwd myuser
- exit
- lxc info my-centos|grep eth0 #write down your IP addr, e.g., 10.169.173.239
- ssh -X myuser@10.169.173.239
- xeyes #should display on your desktop
- exit
VirtualBox inside LXD (requires X11, see above):
- lxc exec my-centos -- /bin/bash
- cd /etc/yum.repos.d
- wget http://download.virtualbox.org/virtualbox/rpm/rhel/virtualbox.repo
- #edit virtualbox.repo
- repo_gpgcheck=0
- yum install VirtualBox-5.0
- useradd myuser
- passwd myuser
- usermod -a -G vboxusers myuser
- exit
- lxc info my-centos|grep eth0 #write down your IP addr, e.g., 10.169.173.239
- ssh -X myuser@10.169.173.239
- VirtualBox #should display on your desktop
- exit
Docker inside of LXD:
- lxc launch ubuntu-daily:16.04 docker
- lxc exec docker -- apt update
- lxc exec docker -- apt dist-upgrade -y
- lxc exec docker -- apt install docker.io -y
- lxc exec docker -- docker run --detach --name app carinamarina/hello-world-app
- lxc exec docker -- docker run --detach --name web --link app:helloapp -p 80:5000 carinamarina/hello-world-web
- lxc list #copy IP for eth0, say 10.178.150.73
- curl http://10.178.150.73 #output: The linked container said... "Hello World!"
- lxc stop docker
- lxc delete docker #if you don't need it anymore

BeeGFS
- cd /mnt/beegfs #to access it from compute nodes
- mkdir $USER #to create your subdirectory (only needs to be done once)
- chmod 700 $USER #to ensure others cannot access you data (only done once)
- cd $USER #go to directory where you should place your large files
- about 160TB of storage over 16 servers (10TB each)
- Currently limited by 1Gpbs switch connection (eth0)
- Not protected by RAID, not backed up!

PAPI
- module load papi
- Reads hardware performance counters
- Check supported counters: papi_avail
- Edit your source file to define performance counter events,
read them and then print or process them, see
PAPI API
- Add to the Makefile compile options: -I${PAPI_INC}
- Add to the Makefile linker options: -L${PAPI_LIB} -lpapi

likwid V5.2.0
- Pins threads to specific cores, avoids
Linux-based thread migration and may increase NUMA performance,
see likwid
project for a complete list of tools (power, pinning etc.)
- print NUMA core topology: likwid-topology -c -g
- Use likwid-pin
to pin threads to specific cores
- Example: likwid-pin myapp
- Example: mpirun -np 2 /usr/local/bin/likwid-pin ./myapp
- Use likwid-perfctr
or likwid-mpirun mearure performance counters, optionally with pinned threads
- Others: likwid-mpirun, likwid-powermeter, likwid-setfreq, ...

Hadoop Map-Reduce and Spark
Simple setup of multi-node
Hadoop map-reduce with HDFS, see
also free
AWS setup as an alternative and the original single
node and
cluster
setup. But follow the instructions below for ARC. Other
components, e.g., YARN, can be added to the setup below as well (not
covered). We'll set up a Hadoop instance with nodes cXXX and cYYY
(optionally more), so you should have gotten at least 2 nodes with srun.
#append to your ~/.bashrc:
module load java
#then issue the command from a shell:
module load java
#distr config, subsitute MY-UNITY-ID with your login ID
mkdir hadoop
cd hadoop
mkdir -p etc/hadoop
cd etc/hadoop
#create file core-site.xml
<configuration>
<property>
<name>fs.defaultFS</name>
<value>hdfs://cXXX:9000</value>
</property>
</configuration>
#create file hdfs-site.xml
<configuration>
<property>
<name>dfs.replication</name>
<value>1</value>
</property>
<property>
<name>dfs.datanode.data.dir</name>
<value>/tmp/MY-UNITY-ID/name/data</value>
</property>
<property>
<name>dfs.namenode.name.dir</name>
<value>/tmp/MY-UNITY-ID/name</value>
</property>
</configuration>
#create mapred-site.xml
<configuration>
<property>
<name>mapred.job.tracker</name>
<value>cXXX:9001</value>
</property>
</configuration>
#create file masters
cXXX
#create file slaves
cXXX
cYYY
#etc.
#for each cXXX/Y/..., create directories
ssh cXXX rm fr /tmp/MY-UNITY-ID
ssh cXXX mkdir -p /tmp/MY-UNITY-ID
ssh cYYY rm fr /tmp/MY-UNITY-ID
ssh cYYY mkdir -p /tmp/MY-UNITY-ID
...
cd ../..
mkdir bin
cd bin
ln -s /usr/local/hadoop/bin/* .
cd ..
mkdir libexec
cd libexec
ln -s /usr/local/hadoop/libexec/* .
cd ..
mkdir sbin
cd sbin
ln -s /usr/local/hadoop/sbin/* .
cd ..
ln -s /usr/local/hadoop/* .
export HADOOP_HOME=`pwd`
export HADOOP_INSTALL=$HADOOP_HOME
export HADOOP_COMMON_LIB_NATIVE_DIR=$HADOOP_INSTALL/lib/native
export HADOOP_OPTS="-Djava.library.path=$HADOOP_INSTALL/lib/native"
export PATH="$PATH:$HADOOP_HOME/bin"
export CLASSPATH=$CLASSPATH:`hadoop classpath`
#distr test: You will get warnings and ssh errors for some command, igore them for now
hdfs getconf -namenodes
hdfs namenode -format
sbin/start-dfs.sh
hdfs dfs -mkdir /user
hdfs dfs -mkdir /user/MY-UNITY-ID
hdfs dfs -put /usr/local/hadoop/etc/hadoop /user/MY-UNITY-ID/input
hadoop jar share/hadoop/mapreduce/hadoop-mapreduce-examples-2.7.2.jar grep /user/MY-UNITY-ID/input /user/MY-UNITY-ID/output 'dfs[a-z.]+'
hdfs dfs -get /user/MY-UNITY-ID/output output
cat output/*
sbin/stop-dfs.sh
 To get rid of ssh errors, you need to add a secondary node server and
other optional services. This is not required, it's an option.
To get rid of ssh errors, you need to add a secondary node server and
other optional services. This is not required, it's an option.
You can also run Spark on top of
Hadoop as follows, which will also default to the HDFS file system:
export SPARK_DIST_CLASSPATH=$(hadoop classpath)
export SPARK_HOME=/usr/local/spark
export CLASSPATH="$CLASSPATH:$SPARK_HOME/lib/*"
export PATH="$PATH:$SPARK_HOME/bin"
run-example SparkPi 10

Tensorflow (2.4)
- Tensorflow
- Notice: Do not pip install your own tensorflow, it will not work!
Same for keras, use tensorflow.keras instead (already installed).
- module load cuda
- python3
import tensorflow as tf
msg = tf.constant('TensorFlow 2.0 Hello World')
tf.print(msg)
- python3 -m pip3 install --upgrade pip #to upgrade pip
- export PYTHONPATH=$PYTHONPATH:$HOME/.local #to include user local packages
- pip3 install --user pkg-name #to install other python packages as user
- python3 setup.py [install] --user #to install python packages as user via setup scripts
- Tensorflow 1.12 with python2 (legacy, being phased out):
- export LD_LIBRARY_PATH=/usr/local/cuda-8.0/lib64:$LD_LIBRARY_PATH
- python2
import tensorflow as tf
hello = tf.constant('Hello, TensorFlow!')
sess = tf.Session()
print(sess.run(hello))
- Also available: python/python2 (version 2.7), pip/pip2: use same user install procedure as above
- Also available: python3.4 (version 3.4), pip3.4: use same user install procedure as above
- Also available: python3 (version 3.6), pip3: use same user install procedure as above
- Also available: python3.8 (version 3.8), pip3.8: use same user install procedure as above, notice the path though: /usr/local/bin/python3.8
- for jupyter-notebook to work, issue
pip3 install jupyter seaborn pydot pydotplus graphviz -U --user
jupyter-notebook --NotebookApp.token='' --ip=cXX *.ipynb
#from your VPN/campus machine, assuming a port 8888 in the printed URL, issue:
ssh arc.csc.ncsu.edu -L 8889:cXX:8888
#point your local browser at https://localhost:8889

PyTorch

Other Packges
A number of packages have been installed, please check out their
location (via: rpm -ql pkg-name) and documentation (see URLs) in this
PDF if you need them. (Notice, only the mvapich2/openmpi/gnu variants
are installed.) Typically, you can get access to them via:
module avail # show which modules are available
module load X
export |grep X #shows what has been defined
gcc/mpicc -I${X_INC} -L{X_LIB} -lx #for a library
./X #for a tool/program, may be some variant of 'X' depending on toolkit
module switch X Y #for mutually exclusive modules if X is already loaded
module unload X
module info #learn how to use modules
Current list of available modules:
-------------------- /opt/ohpc/pub/moduledeps/gnu-mvapich2 ---------------------
adios/1.10.0 mpiP/3.4.1 petsc/3.7.0 scorep/3.0
boost/1.61.0 mumps/5.0.2 phdf5/1.8.17 sionlib/1.7.0
fftw/3.3.4 netcdf/4.4.1 scalapack/2.0.2 superlu_dist/4.2
hypre/2.10.1 netcdf-cxx/4.2.1 scalasca/2.3.1 tau/2.26
imb/4.1 netcdf-fortran/4.4.4 scipy/0.18.0 trilinos/12.6.4
------------------------- /opt/ohpc/pub/moduledeps/gnu -------------------------
R_base/3.3.1 metis/5.1.0 ocr/1.0.1 pdtoolkit/3.22
gsl/2.2.1 mvapich2/2.2 (L) openblas/0.2.19 superlu/5.2.1
hdf5/1.8.17 numpy/1.11.1 openmpi/1.10.4
------------------------- /opt/ohpc/admin/modulefiles --------------------------
spack/0.8.17
-------------------------- /opt/ohpc/pub/modulefiles ---------------------------
EasyBuild/2.9.0 java pgi-llvm
autotools (L) ohpc (L) pgi-nollvm
cuda (L) openmpi3/3.1.4 prun/1.1
gnu/5.4.0 (L) papi/5.4.3 prun/1.3 (L,D)
gnu8/8.3.0 pgi/19.10 valgrind/3.11.0

Advanced topics (pending)
For all other topics, access is restricted. Request a root password.
Also, read this documentation, which is only
accessible from selected NCSU labs.
This applies to:
- booting your own kernel
- installing your own OS

Known Problems
Consult the FAQ. If this does not help, then
please report your problem.
References:
- A User's
Guide to MPI by Peter Pacheco
- Debugging: Gdb only works on one task with MPI, you need to
"attach" to other tasks on the respective nodes. We don't have
totalview (an MPI-aware debugger). You can also use
printf debugging, of course. If your program SEGVs, you can
set ulimit -c unlimited and run the mpi program again, which
will create one or more core dump files (per rank)
named "core.PID", which you can then debug: gdb binary and
then core core.PID.

Additional references:





























 To get rid of ssh errors, you need to add a secondary node server and
other optional services. This is not required, it's an option.
To get rid of ssh errors, you need to add a secondary node server and
other optional services. This is not required, it's an option.




